Apolipoprotein E gene polymorphism influenced glycemic status among Malaysians
DOI:
https://doi.org/10.15419/bmrat.v6i7.557Keywords:
APOE, FBG, polymorphism, T2DMAbstract
Aim: In the last decade Apolipoprotein E (APOE) gene polymorphism has been identified as one of the risk factors of type 2 diabetes mellitus (T2DM). Though more than 11% population of Malaysia are suffering from T2DM, there is inadequate data on the correlation between the APOE gene polymorphism and pathogenesis of diabetes among Malaysians. Hence, in this study we aimed to find out the association between the frequencies of APOE allele and fasting blood glucose (FBG) concentration among subjects with T2DM.
Methods: A total of 102 subjects were recruited into two distinct groups, 51 in diabetes (cases) and 51 in non-diabetes (control) group. Their fasting blood sample was tested for FBG, while APOE genotyping was carried out using restriction fragment length polymorphism technique. Predictive Analytics Software (PASW) statistics, version 18.0, was used for statistical analyses.
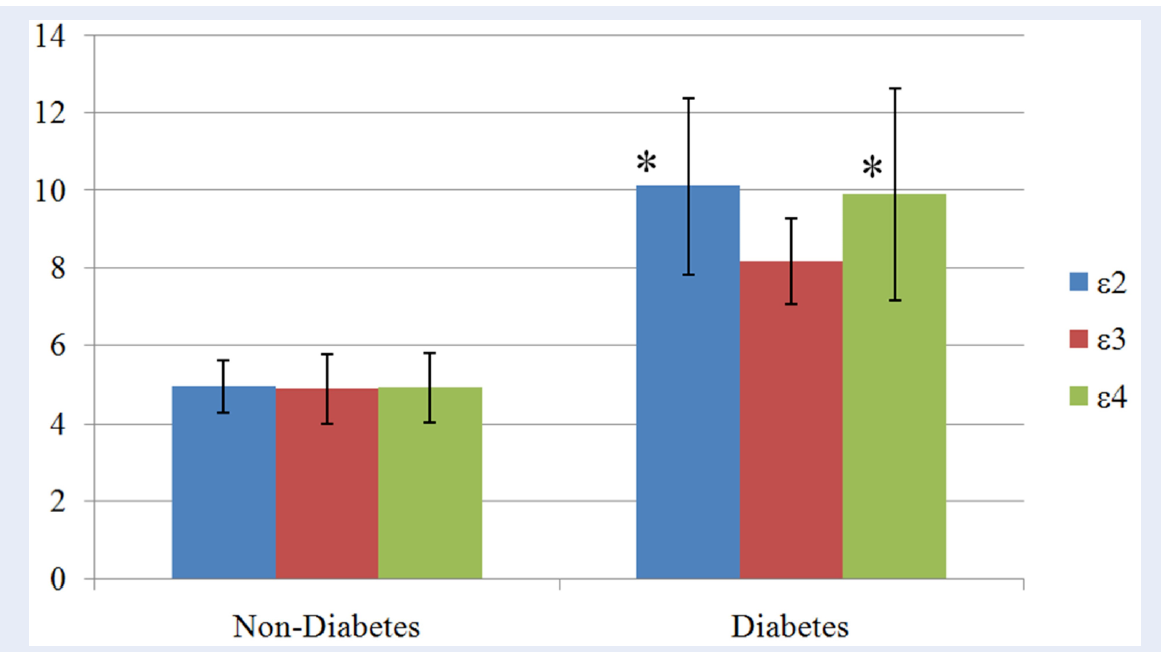
Results: There was no association between APOE alleles and T2DM; odd ratios for the e2, e3 and e4 alleles were 1.51 (95%CI: 0.615-3.706), 0.77 (95%CI: 0.431-1.375) and 1.12 (95%CI: 0.584-2.131) respectively. The highest mean FBG was found in subjects with e2 alleles, followed by e4 and e3 alleles in both cases and control groups. Both e2 and e4 alleles were significantly linked to higher mean FBG (p=0.03 and 0.04 for the respectively) compared to e3 allele in diabetes group.
Conclusions: Although the APOE gene was not found to be associated with T2DM, it may influence glycemic status among subjects.
Downloads
Published
Issue
Section
License
Copyright The Author(s) 2017. This article is published with open access by BioMedPress. This article is distributed under the terms of the Creative Commons Attribution License (CC-BY 4.0) which permits any use, distribution, and reproduction in any medium, provided the original author(s) and the source are credited.
